Interoperability with web-applications
The mdatools package works nicely together with interactive web-applications available at https://mda.tools. There is a dedicated app, https://mda.tools/prep, which lets you create a preprocessing chain, train its parameters and apply it to new data. If you open the app and play with it you will notice that it contains all preprocessing methods mentioned above.
Any preprocessing model created using prep.fit() like in the example shown in the previous section can be saved into a JSON file and then be loaded into the Apply tab of the web-application.
Here is an example:
# define calibration set
Xc <- simdata$spectra.c
w <- simdata$wavelength
attr(Xc, "xaxis.values") <- w
attr(Xc, "xaxis.name") <- "Wavelength, nm"
# create a sequence of preprocessing methods
myprep <- list(
prep("savgol", width = 9, porder = 1),
prep("emsc", degree = 0),
prep("norm", type = "area"),
prep("varsel", var.ind = w < 300),
prep("center")
)
# train the preprocessing model
myprep <- prep.fit(myprep, Xc)
# save the model to JSON file
writeJSON(myprep, "prep-model.json")If you run the code on your computer you will see a new prep-model.json file created in the working directory. Now open https://mda.tools/prep in your browser, select tab Apply and load the file with the preprocessing model. You will see something similar to the following:
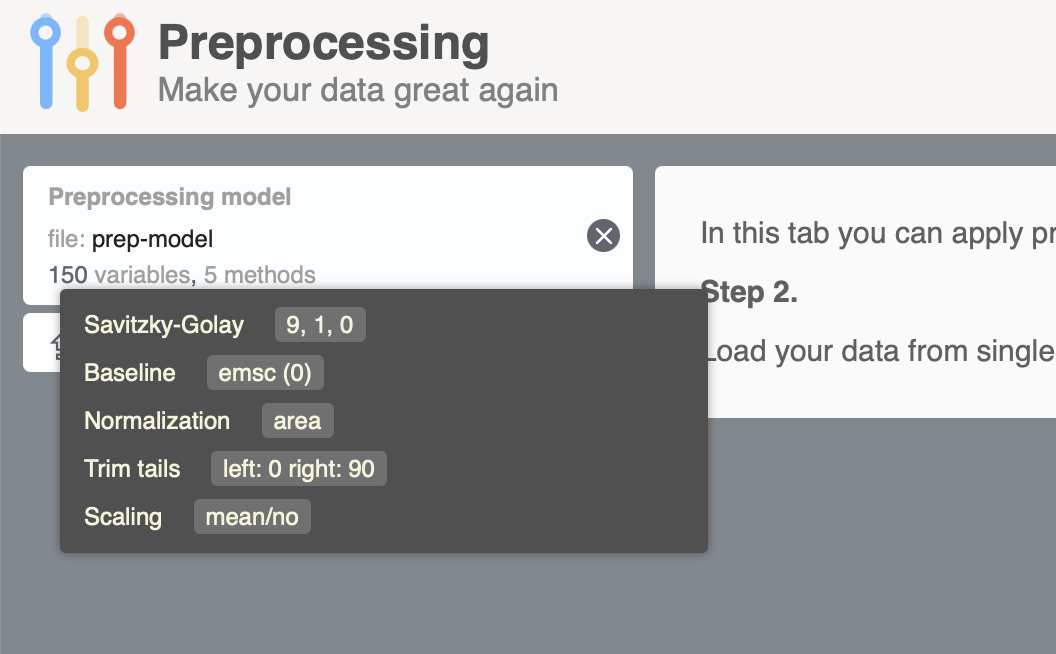
After that you can upload dataset and preprocess it in the app.
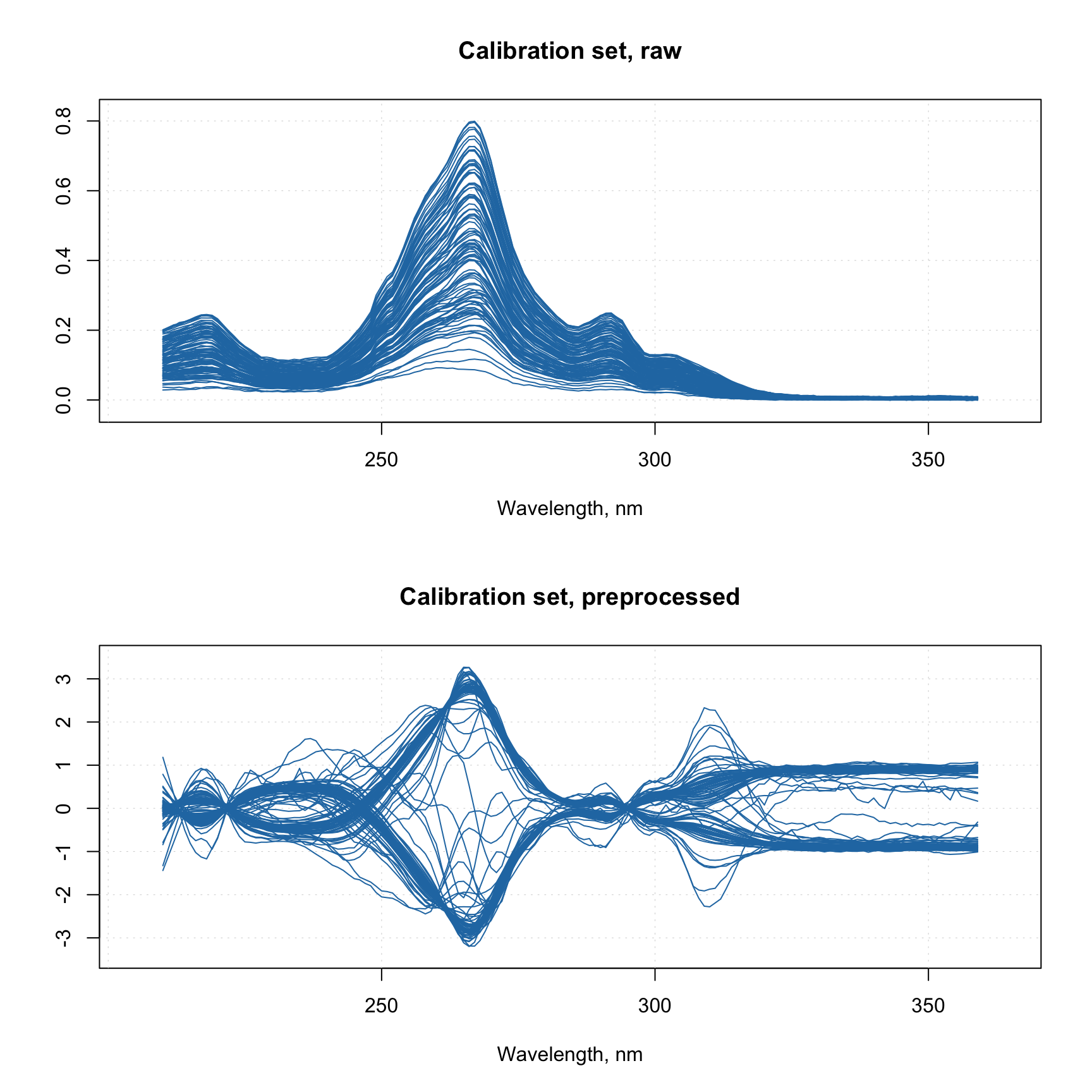
Likewise, any preprocessing model created in the application can be saved into a JSON file and loaded into R, so you can use it via prep.apply(). For example this JSON file I created using the Train tab of the app and Simdata. It does SG smoothing, SNV normalization, and mean centering. Download it to your working directory and run the following code (here I assume you already have Xc matrix from the previous block of code):
# load preprocessing model from JSON file
myprep <- readJSON("prep-model-fromweb.json")
# apply it to Xc
Xcp <- prep.apply(myprep, Xc)
# show the result
par(mfrow = c(2, 1))
mdaplot(Xc, type = "l", main = "Calibration set, raw")
mdaplot(Xcp, type = "l", main = "Calibration set, preprocessed")