Variable selection as preprocessing method
Variable selection can be done by using mda.exclcols(), which simply hides variables/columns, which must not be taken into account in calculations, or by mda.subset() which selects only desired columns and removes the rest. Both methods preserve all additional attributes assigned to the data.
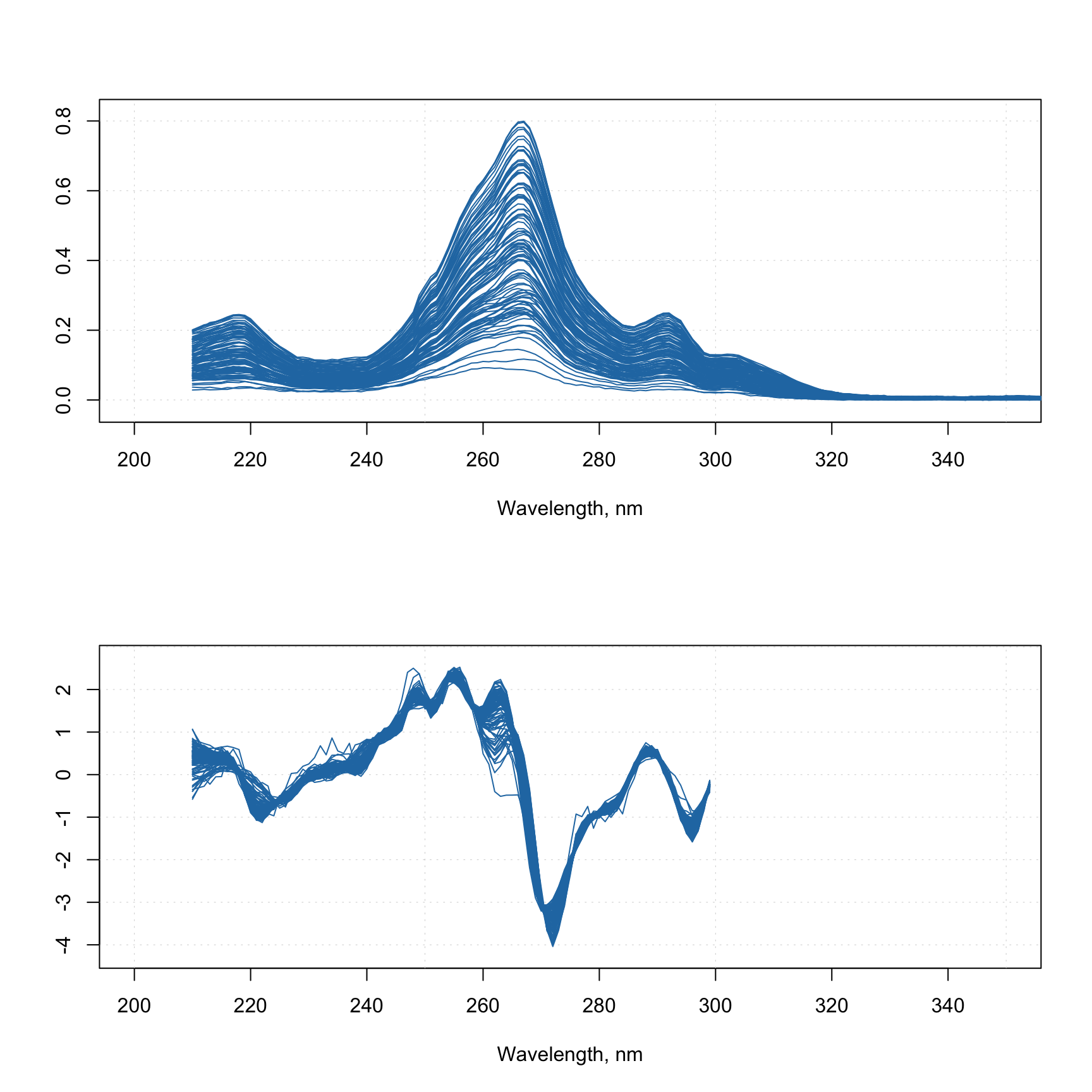
The method prep.varsel() is simply a wrapper, which allows selection of only desired variables (similar to mda.subset()) but can also be incorporated into preprocessing workflow (see next section for details). In the example below it is used to trim tails of spectra.
# load spectra from the Simdata and add some attributes
data(simdata)
X <- simdata$spectra.c
w <- simdata$wavelength
attr(X, "xaxis.values") <- w
attr(X, "xaxis.name") <- "Wavelength, nm"
attr(X, "name") <- "Simdata"
# apply variable selection as preprocessing
col.ind = w > 240 & w < 300
Y <- prep.varsel(X, col.ind)
# show both original and preprocessed spectra
par(mfrow = c(2, 1))
mdaplot(X, type = "l", xlim = c(200, 380))
mdaplot(Y, type = "l", xlim = c(200, 380))